-Search query
-Search result
Showing 1 - 50 of 88 items for (author: takada & k)

EMDB-35029: 
SARS-CoV2 spike protein with ACE2, no ACE2 binding.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35030: 
SARS-CoV2 spike protein with ACE2. 1 ACE2 bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35031: 
SARS-CoV2 spike protein with ACE2. 2 ACE2 bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35032: 
SARS-CoV2 spike protein with ACE2. 3 ACE2 bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35036: 
SARS-CoV2 spike protein with ACE2 decoy.no ACE2 decoy binding
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35037: 
SARS-CoV2 spike protein with ACE2 decoy. 1 ACE2 decoy bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35038: 
SARS-CoV2 spike protein with ACE2 decoy. 1 ACE2 decoy bound and 2 RBD up form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35039: 
SARS-CoV2 spike protein with ACE2 decoy. 2 ACE2 decoy bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35040: 
SARS-CoV2 spike protein with ACE2 decoy. 3 ACE2 decoy bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-36345: 
RBD of SARS-CoV2 spike protein with ACE2 decoy
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

PDB-8jje: 
RBD of SARS-CoV2 spike protein with ACE2 decoy
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T
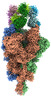
EMDB-33972: 
Cryo-EM structure of the SARS-CoV-2 spike protein (3-up RBD) bound to neutralizing antibody CSW1-1805
Method: single particle / : Anzai I, Fujita J, Ono C, Kosaka Y, Miyamoto Y, Shichinohe S, Takada K, Torii S, Taguwa S, Suzuki K, Makino F, Kajita T, Inoue T, Namba K, Watanabe T, Matsuura Y
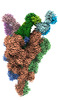
EMDB-33973: 
Cryo-EM structure of the SARS-CoV-2 spike protein (1-up RBD) bound to neutralizing antibody CSW2-1353
Method: single particle / : Anzai I, Fujita J, Ono C, Kosaka Y, Miyamoto Y, Shichinohe S, Takada K, Torii S, Taguwa S, Suzuki K, Makino F, Kajita T, Inoue T, Namba K, Watanabe T, Matsuura Y
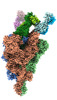
EMDB-33974: 
Cryo-EM structure of the SARS-CoV-2 spike protein (2-up RBD) bound to neutralizing antibody CSW2-1353
Method: single particle / : Anzai I, Fujita J, Ono C, Kosaka Y, Miyamoto Y, Shichinohe S, Takada K, Torii S, Taguwa S, Suzuki K, Makino F, Kajita T, Inoue T, Namba K, Watanabe T, Matsuura Y
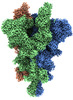
EMDB-33975: 
Cryo-EM structure of the SARS-CoV-2 spike protein (2-up RBD) with C480A mutation
Method: single particle / : Anzai I, Fujita J, Ono C, Kosaka Y, Miyamoto Y, Shichinohe S, Takada K, Torii S, Taguwa S, Suzuki K, Makino F, Kajita T, Inoue T, Namba K, Watanabe T, Matsuura Y

EMDB-27637: 
Structure of EBOV GP lacking the mucin-like domain with 2.1.1D5 scFv and 6D6 scFv bound
Method: single particle / : Yu X, Saphire EO

EMDB-27638: 
Structure of EBOV GP lacking the mucin-like domain with 9.20.1A2 Fab and 6D6 scFv bound
Method: single particle / : Yu X, Saphire EO

PDB-8dpl: 
Structure of EBOV GP lacking the mucin-like domain with 2.1.1D5 scFv and 6D6 scFv bound
Method: single particle / : Yu X, Saphire EO

PDB-8dpm: 
Structure of EBOV GP lacking the mucin-like domain with 9.20.1A2 Fab and 6D6 scFv bound
Method: single particle / : Yu X, Saphire EO

EMDB-16246: 
ARE-ABCF VmlR2 bound to a 70S ribosome
Method: single particle / : Crowe-McAuliffe C, Wilson DN

PDB-8buu: 
ARE-ABCF VmlR2 bound to a 70S ribosome
Method: single particle / : Crowe-McAuliffe C, Wilson DN

EMDB-13242: 
PoxtA-EQ2 antibiotic resistance ABCF bound to E. faecalis 70S ribosome, state I
Method: single particle / : Crowe-McAuliffe C, Wilson DN

PDB-7p7r: 
PoxtA-EQ2 antibiotic resistance ABCF bound to E. faecalis 70S ribosome, state I
Method: single particle / : Crowe-McAuliffe C, Wilson DN

EMDB-13241: 
E. faecalis 70S ribosome bound by PoxtA-EQ2, high-resolution combined volume
Method: single particle / : Crowe-McAuliffe C, Wilson DN

EMDB-13243: 
PoxtA-EQ2 antibiotic resistance ABCF bound to E. faecalis 70S ribosome, state II
Method: single particle / : Crowe-McAuliffe C, Wilson DN

EMDB-13244: 
PoxtA-EQ2 antibiotic resistance ABCF bound to E. faecalis 70S ribosome, state III
Method: single particle / : Crowe-McAuliffe C, Wilson DN

EMDB-13245: 
E. faecalis 70S ribosome with P-tRNA, state IV
Method: single particle / : Crowe-McAuliffe C, Wilson DN

PDB-7p7q: 
E. faecalis 70S ribosome bound by PoxtA-EQ2, high-resolution combined volume
Method: single particle / : Crowe-McAuliffe C, Wilson DN

PDB-7p7s: 
PoxtA-EQ2 antibiotic resistance ABCF bound to E. faecalis 70S ribosome, state II
Method: single particle / : Crowe-McAuliffe C, Wilson DN

PDB-7p7t: 
PoxtA-EQ2 antibiotic resistance ABCF bound to E. faecalis 70S ribosome, state III
Method: single particle / : Crowe-McAuliffe C, Wilson DN

PDB-7p7u: 
E. faecalis 70S ribosome with P-tRNA, state IV
Method: single particle / : Crowe-McAuliffe C, Wilson DN

EMDB-13017: 
RqcH DR variant bound to 50S-peptidyl-tRNA-RqcP RQC complex (rigid body refinement)
Method: single particle / : Crowe-McAuliffe C, Wilson DN

PDB-7ope: 
RqcH DR variant bound to 50S-peptidyl-tRNA-RqcP RQC complex (rigid body refinement)
Method: single particle / : Crowe-McAuliffe C, Wilson DN

EMDB-12331: 
LsaA, an antibiotic resistance ABCF, in complex with 70S ribosome from Enterococcus faecalis
Method: single particle / : Crowe-McAuliffe C, Kasari M, Hauryliuk VH, Wilson DN

EMDB-12332: 
VgaA-LC, an antibiotic resistance ABCF, in complex with 70S ribosome from Staphylococcus aureus
Method: single particle / : Crowe-McAuliffe C, Murina V

EMDB-12333: 
70S ribosome from Staphylococcus aureus
Method: single particle / : Crowe-McAuliffe C, Murina V

EMDB-12334: 
VgaL, an antibiotic resistance ABCF, in complex with 70S ribosome from Listeria monocytogenes
Method: single particle / : Crowe-McAuliffe C, Turnbull KJ

PDB-7nhk: 
LsaA, an antibiotic resistance ABCF, in complex with 70S ribosome from Enterococcus faecalis
Method: single particle / : Crowe-McAuliffe C, Kasari M, Hauryliuk VH, Wilson DN

PDB-7nhl: 
VgaA-LC, an antibiotic resistance ABCF, in complex with 70S ribosome from Staphylococcus aureus
Method: single particle / : Crowe-McAuliffe C, Murina V, Hauryliuk V, Wilson DN

PDB-7nhm: 
70S ribosome from Staphylococcus aureus
Method: single particle / : Crowe-McAuliffe C, Murina V, Hauryliuk V, Wilson DN

PDB-7nhn: 
VgaL, an antibiotic resistance ABCF, in complex with 70S ribosome from Listeria monocytogenes
Method: single particle / : Crowe-McAuliffe C, Turnbull KJ, Hauryliuk V, Wilson DN

EMDB-11890: 
Bacillus subtilis ribosome-associated quality control complex state A. Ribosomal 50S subunit with peptidyl tRNA in the A/P position and RqcH.
Method: single particle / : Crowe-McAuliffe C, Wilson DN

EMDB-11891: 
Bacillus subtilis ribosome-associated quality control complex state B, multibody refinement focussed on RqcH. Ribosomal 50S subunit with P-tRNA, RqcH, and RqcP/YabO
Method: single particle / : Crowe-McAuliffe C, Wilson DN

PDB-7as9: 
Bacillus subtilis ribosome-associated quality control complex state A. Ribosomal 50S subunit with peptidyl tRNA in the A/P position and RqcH.
Method: single particle / : Crowe-McAuliffe C, Wilson DN

PDB-7asa: 
Bacillus subtilis ribosome-associated quality control complex state B, multibody refinement focussed on RqcH. Ribosomal 50S subunit with P-tRNA, RqcH, and RqcP/YabO
Method: single particle / : Crowe-McAuliffe C, Wilson DN

EMDB-11889: 
Bacillus subtilis ribosome quality control complex state B. Ribosomal 50S subunit with P-tRNA, RqcH, and RqcP/YabO
Method: single particle / : Crowe-McAuliffe C, Wilson DN

EMDB-11913: 
Bacillus subtilis ribosome-associated quality control complex, state C, derived from RqcH affinity purification.
Method: single particle / : Crowe-McAuliffe C, Wilson DN

EMDB-11914: 
Ribosome-associated quality control complex from Bacillus subtilis, state D. Large ribosomal subunit in complex with P-tRNA and RqcP/YabO.
Method: single particle / : Crowe-McAuliffe C, Wilson DN

EMDB-11915: 
Bacillus subtilis ribosome-associated quality control complex state B*, derived from RqcH affinity purification.
Method: single particle / : Crowe-McAuliffe C, Wilson DN

EMDB-11916: 
Bacillus subtilis ribosome-associated quality control complex, state E, derived from RqcP/YabO affinity purification.
Method: single particle / : Crowe-McAuliffe C, Wilson DN
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
